Department of Connectomics
Moritz Helmstaedter
Our goal is to decipher how the cerebral cortex stores sensory experience and uses it to detect objects in the current environment.
To this end, we develop and apply methods for measuring communication maps of neuronal circuits, connectomes. We strive to push the frontiers of connectomics to make the mapping of neuronal circuits a high-throughput technique.
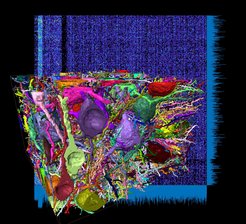
First connectome from the mammalian cerebral cortex: connectivity matrix between thousands of axons and dendrites in a piece of layer 4 from the somatosensory cortex of a mouse. While about 4 times larger than the 2013 retina connectome, the reconstruction only required about 4,000 work hours by student annotators, the rest of the work was taken over by improved artificial intelligence.
We want to:
- measure the similarity between neuronal networks in the cortex of different individuals and different species in search for the algorithms of sensory perception
- search for engrams of sensory experience in the cerebral cortex
- understand the alterations in neuronal network structure in psychiatric disease.
To this end, we apply 3-dimensional electron microscopy to scan the complete neuronal circuits in pieces of brain tissue. Key methodological steps are: tissue preparation and staining, electron microscopic imaging, and large-scale data analysis. Our lab is providing state-of-the-art methodology for tissue preparation and data analysis.
How are algorithms implemented in the brain? In contrast to man-made computers, nature-built brains do not distinguish between hard- and software. The most plausible way to store sensory expectations and to implement algorithms in neuronal networks is by shaping their extremely complex connectivity structure. Each of the 17 billion neurons in our cerebral cortices connects directly and specifically to about 1000 other neurons. Imagine! Do you have 1000 friends you regularly talk to one-on-one? Or even facebook-friends?
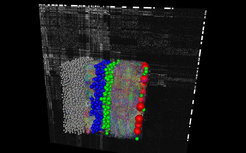
The complexity of these neuronal networks is most plausibly the structural correlate of the complex computational methods implemented in vertebrate brains. But no one has yet been able to measure these networks at high resolution and completeness in the cerebral cortex. How can we then know whether and how the algorithms we are looking for are implemented?
Only since 2000, imaging and analysis methods have been developed that start to allow us to map neuronal networks in completion and increasing throughput. Because neurons are small, electron microscopy is required, but in totally new magnitude.
Analysis of our image data is so difficult that so far, only humans could solve the most difficult problems. So we started by combining computer power with human brain power to reconstruct the neuronal networks of the cerebral cortex. With major improvements in automated data analysis, we are getting closer to a full automation of connectomic data analysis, and some connectomic results are already achievable without any human data annotation.
Cerebral Cortex Connectomics: Background
How does our brain distinguish, say, an apple from a car?
Some expectation of apples and cars must be combined with novel input from our sensors (eyes, ears, touch receptors). For all we know, the sensory expectation of apples and cars (and certainly friends’ faces) is not already encoded in our genomes, but is imprinted into our brains during our lives. We want to understand how in the cerebral cortex, the main part of vertebrates’ brains, such a combination of acquired sensory expectation with sensory novelty is implemented for the detection and classification of objects and events.
This problem is a major scientific problem that challenges us in at least two ways: Not only has neurobiology over the last 100 years not been able to decipher the way the cerebral cortex classifies sensory objects; even more stunningly in spite of decades of artificial intelligence research (cybernetics, machine learning), we have not discovered algorithms that are even close to as general and powerful. No computer today is even close to a vertebrate brain in its ability to classify people in a video stream or hidden faces in a visual scene.
It is therefore our ambition to decipher the algorithms that are implemented in vertebrate brains for complex pattern recognition.





